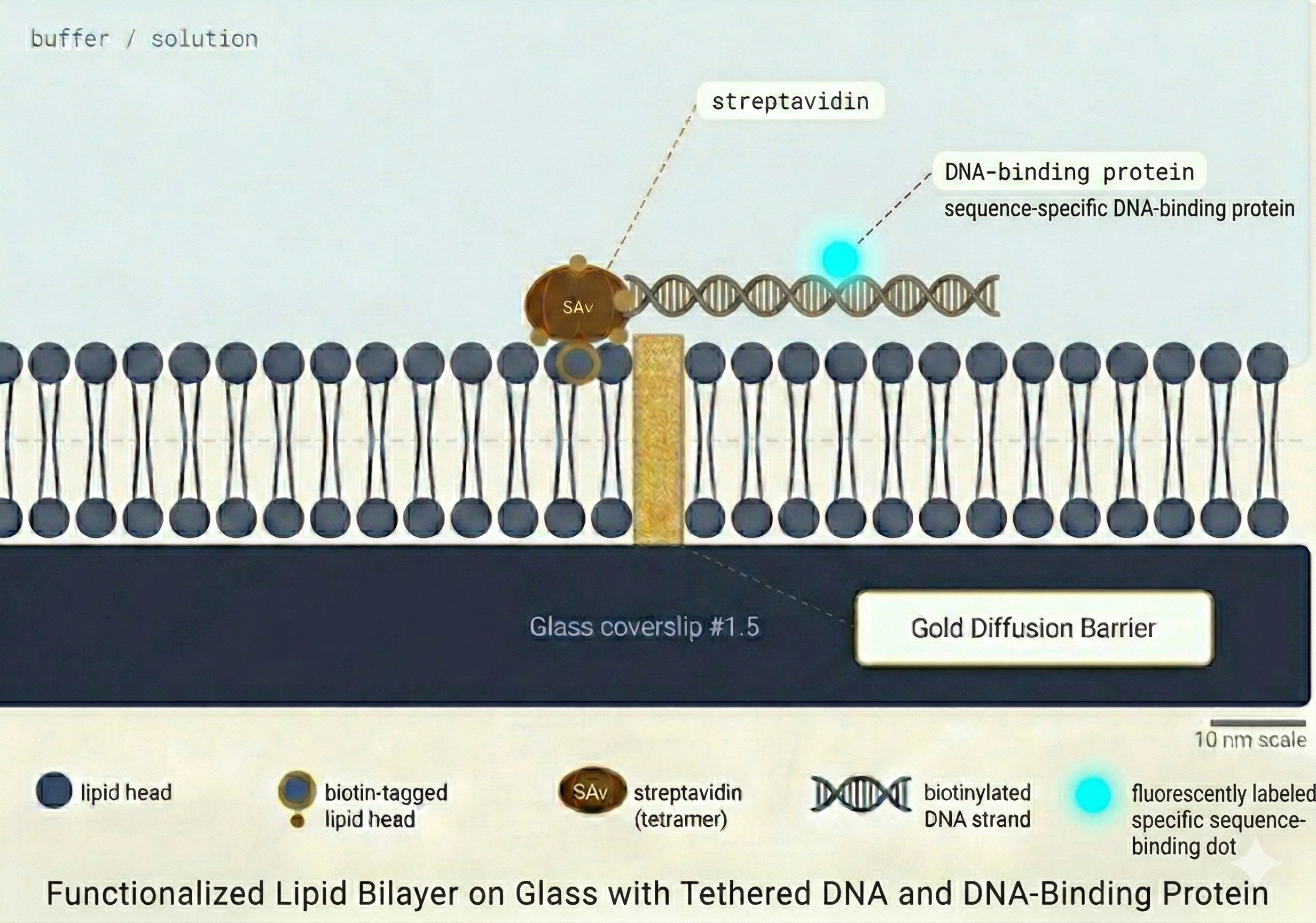
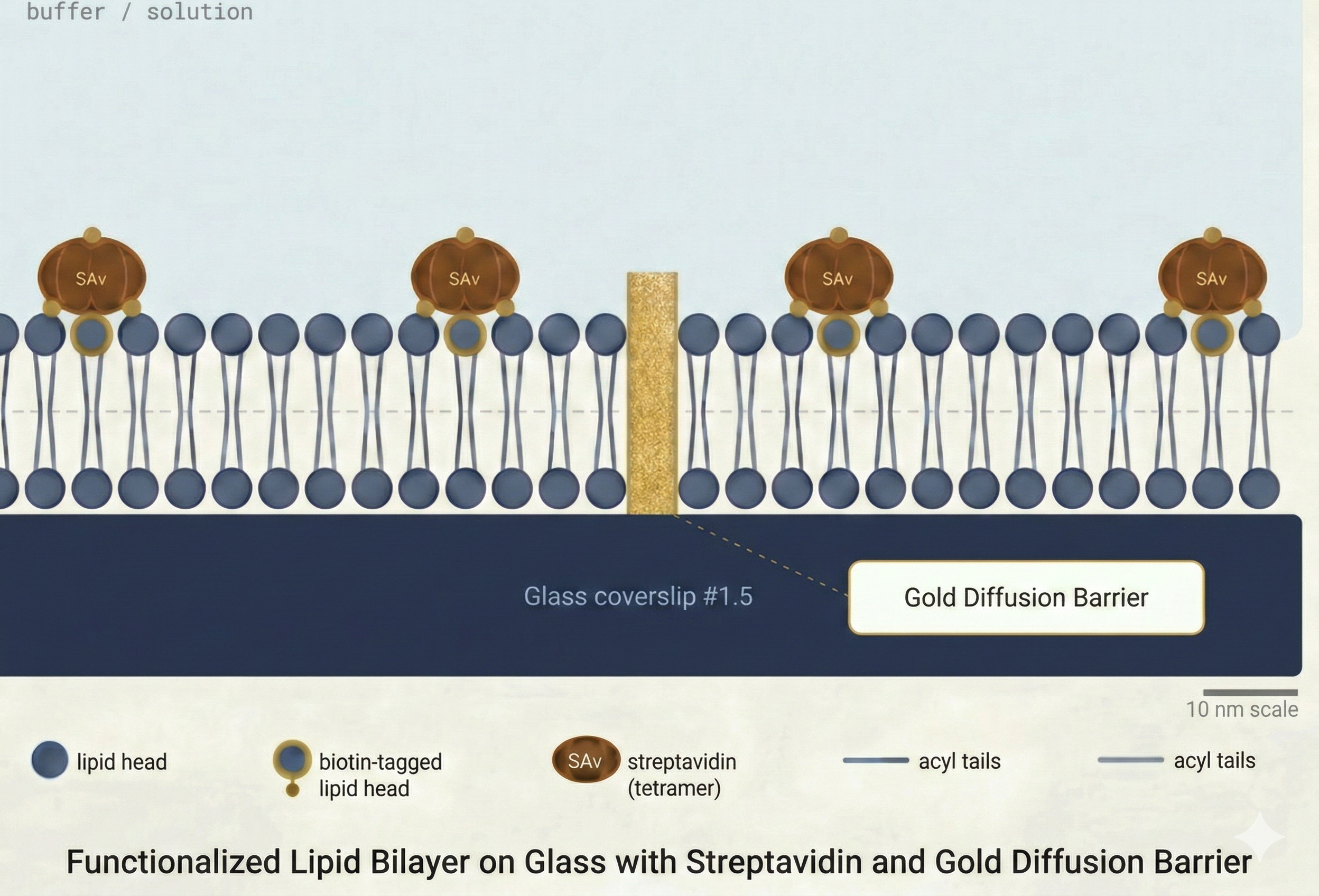
Lipid Bilayer Formation
Before any DNA experiment, all surfaces of the microfluidic flow channel are passivated using a supported lipid bilayer (SLB). The SLB serves two critical functions:
- Mobility: Lipid-anchored DNA molecules can diffuse along the bilayer surface, allowing them to migrate toward barriers under flow.
- Passivation: The SLB minimizes non-specific adsorption of proteins, reducing background fluorescence and ensuring clean data.
The bilayer is formed directly within the flow cell, using standard vesicle fusion protocols compatible with most lipid compositions.
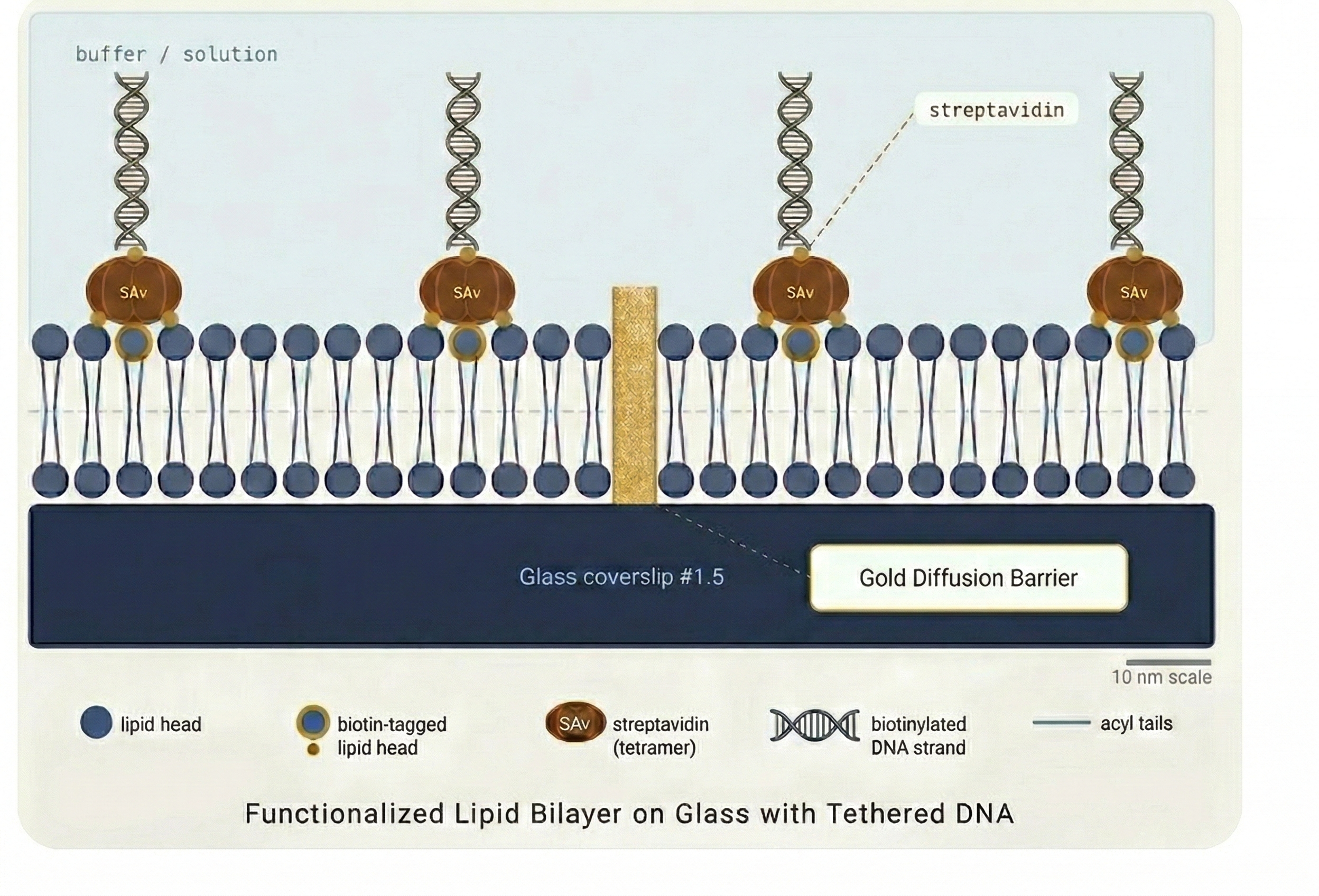
DNA Tethering
DNA molecules (typically 20–50 kbp in length) are tethered to the supported lipid bilayer via a biotin–streptavidin interaction at one end (single tether) or both ends (double tether).
Once introduced into the flow cell, a small and controlled flow is applied. Because the DNA is anchored to the mobile bilayer, the hydrodynamic drag extends and drives the DNA molecules toward the nanofabricated barriers — without stretching or damaging them.
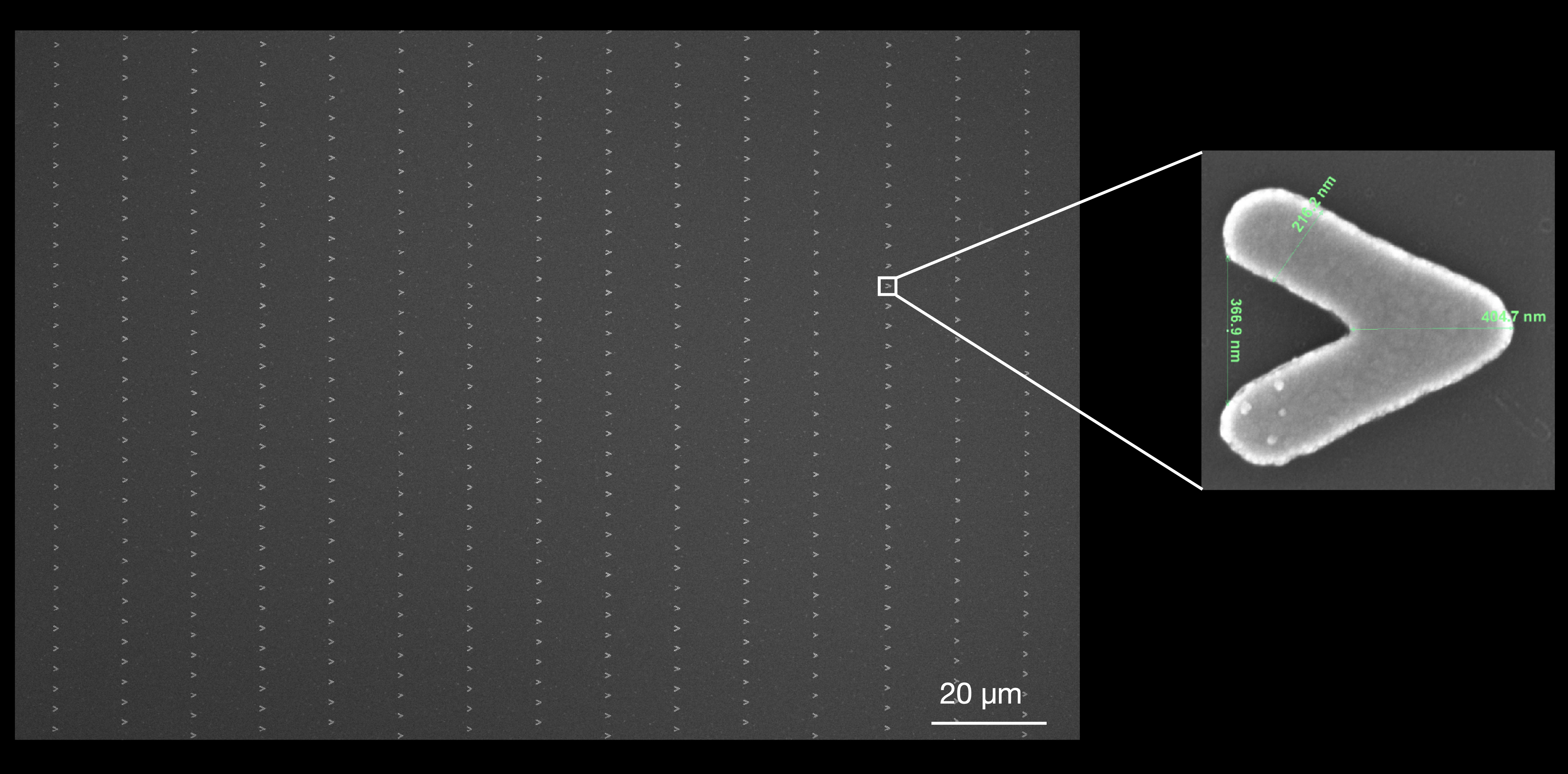
Curtain Formation & Imaging
When DNA molecules reach the nanofabricated barriers, the barriers physically capture them at their leading edges. This forms the DNA Curtain — an ordered, parallel array of individually extended and spatially separated DNA molecules, all aligned along the barrier edge.
With flow maintained, the DNA remain extended and accessible. Fluorescently labeled proteins of interest are then introduced, and their binding, diffusion, and activity along individual DNA molecules is visualized in real time using objective-type TIRF microscopy.
Automated tracking scripts extract kymographs for every molecule in the field of view simultaneously, generating statistically robust, quantitative data on diffusion coefficients, dwell times, processivity, and more — from a single experiment.