DNA Damage Repair
DNA double-strand breaks are among the most cytotoxic lesions a cell can sustain. Understanding how repair complexes recognize, process, and fix these breaks is fundamental to cancer biology, aging research, and genome stability.
DNA Curtains enable direct, real-time observation of repair machinery in action. The MRN complex (Mre11-Rad50-Nbs1), BLM helicase, CtIP, and other key repair factors have been visualized on individual DNA substrates — revealing recruitment dynamics, resection mechanisms, and pathway choice at the single-molecule level.
Fluorescent labeling of specific protein components combined with multi-color TIRF imaging allows simultaneous tracking of multiple repair factors, revealing their spatiotemporal coordination.
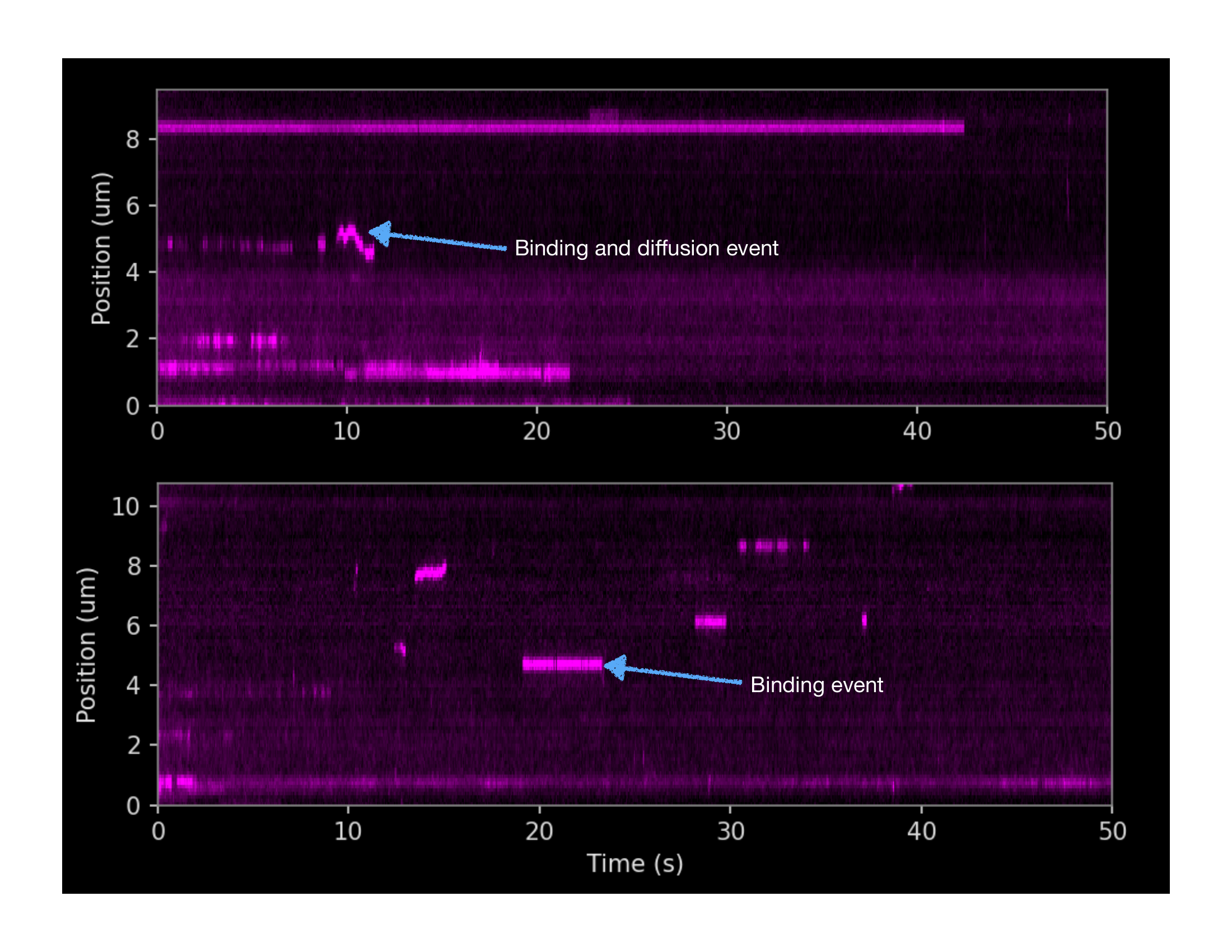
Kymographs
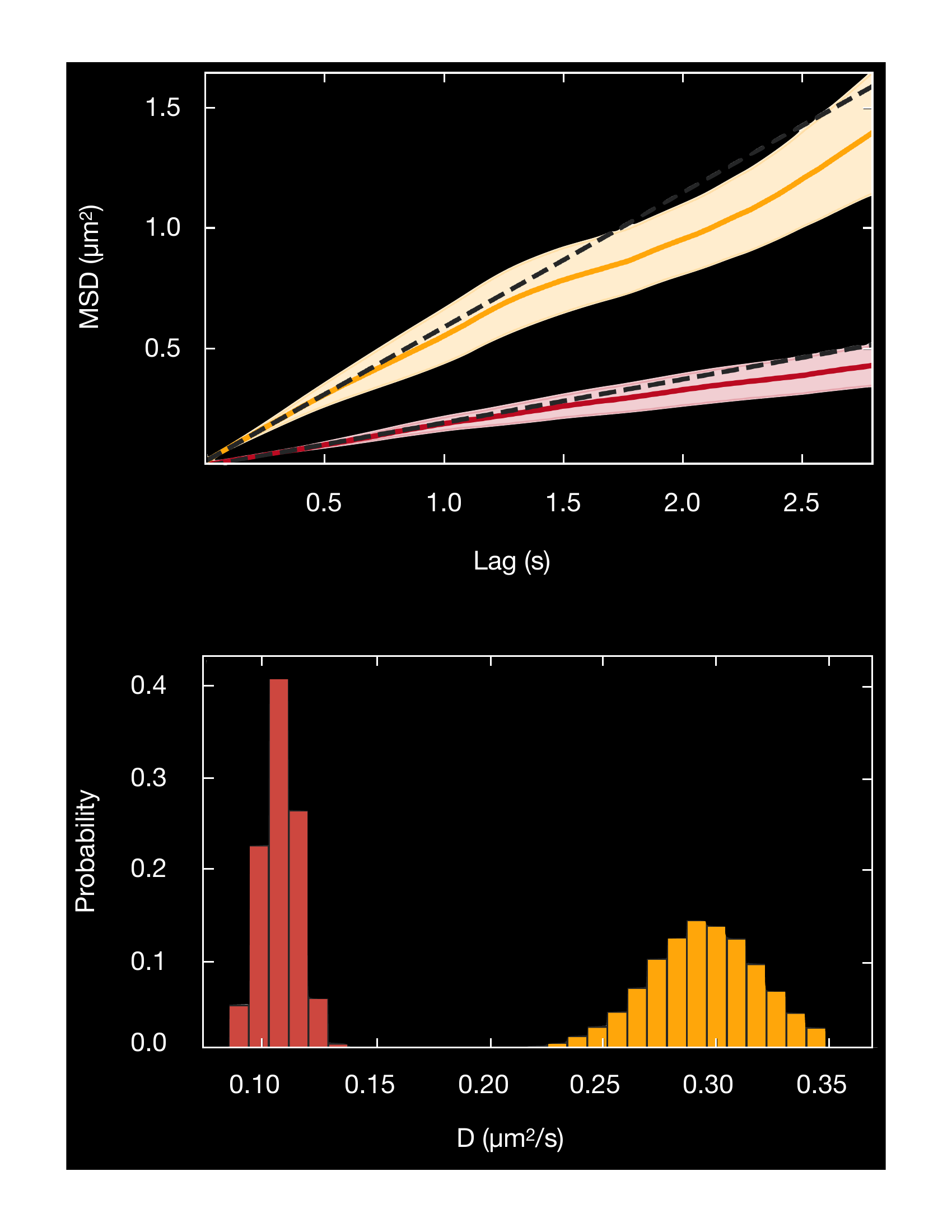
Binding kinetics, and diffusion of DNA-binding proteins are easily visualized as kymographs by picking out an individual DNA tether out of hundreds recorded, and plotting protein binding position along the tether against time.
Both panels on the left show the binding behavior of proteins to one individual DNA tether. Dwell times, diffusion behavior and binding location along the DNA strand can be visualized and quantified.